Robustness
1 Custom functions
rm(list = ls())
fpackage.check <- function(packages) {
lapply(packages, FUN = function(x) {
if (!require(x, character.only = TRUE)) {
install.packages(x, dependencies = TRUE)
library(x, character.only = TRUE)
}
})
}
fsave <- function(x, file = NULL, location = "./data/processed/") {
ifelse(!dir.exists("data"), dir.create("data"), FALSE)
ifelse(!dir.exists("data/processed"), dir.create("data/processed"), FALSE)
if (is.null(file))
file = deparse(substitute(x))
datename <- substr(gsub("[:-]", "", Sys.time()), 1, 8)
totalname <- paste(location, file, "_", datename, ".rda", sep = "")
save(x, file = totalname) #need to fix if file is reloaded as input name, not as x.
}
fload <- function(filename) {
load(filename)
get(ls()[ls() != "filename"])
}
fshowdf <- function(x, ...) {
knitr::kable(x, digits = 3, "html", ...) %>%
kableExtra::kable_styling(bootstrap_options = c("striped", "hover")) %>%
kableExtra::scroll_box(width = "100%", height = "300px")
}
colorize <- function(x, color) {sprintf("<span style='color: %s;'>%s</span>", color, x) }2 Packages
packages = c("ggplot2", "ggpubr", "tidyverse", "RColorBrewer", "stringdist", "stringi", "future.apply",
"gamlss", "gamlss.dist", "gamlss.tr", "splines", "splines2", "psych", "MASS", "dplyr", "flextable",
"fastDummies", "fixest", "ggtext")
# load dplyr last otherwise conflict with select from MASS
fpackage.check(packages)3 Input
We use three processed datasets.
20240717df_ppf.rda: person period file for main analysis
20240717df_pubs.rda: publications for descriptives
20240711linked_pubs_oa_fields.rda: publications linked to fields
df_ppf <- fload(file = "./data/processed/20240717df_ppf.rda")
df_pubs <- fload(file = "./data/processed/20240717df_pubs.rda")
df_fields <- fload(file = "./data/processed/20240711linked_pubs_oa_fields.rda")4 Data wrangling
4.1 Domain
I will use all publications. Also the publications of 2020 and later, as I think this indicates the field/domain the author was working in during Covid.
I don’t pick the top domain, but calculate a percentage within domain. This does imho more justice to multidisciplinary and also takes into account possible arbitrariness in the black box of openalex domain identifier.
domain <- df_fields %>% dplyr::select("domain", "id") %>% #select conflicts with MASS
filter(!is.na(domain)) %>%
mutate(domain2 = as.numeric(as.factor(domain))) %>% #make numeric of domain variable
group_by(id) %>%
mutate(npubs_domain = n(), #number of publications domain is based
health_domain = sum(domain2 == 1) / npubs_domain, #domain as proportions
life_domain = sum(domain2 == 2) / npubs_domain,
physical_domain = sum(domain2 == 3) / npubs_domain,
social_domain = sum(domain2 == 4) / npubs_domain) %>%
ungroup() %>% #only keep one record per id
mutate(id.dup =ifelse(duplicated(id), 1,0)) %>%
filter(id.dup == 0) %>%
dplyr::select("id", "npubs_domain", "health_domain", "life_domain", "physical_domain", "social_domain")
#describe(domain)add to our df_ppf
df_ppf <- df_ppf %>%
left_join(domain)4.2 Gender
df_gender <- df_pubs %>%
ungroup() %>%
mutate(id.dup = ifelse(duplicated(id), 1, 0)) %>%
filter(id.dup == 0) %>%
dplyr::select("id", "gender_2", "firstname", "lastname") %>%
mutate(gender2 = ifelse(is.na(firstname) & is.na(gender_2), 3, gender_2))
df_gender$gender2[is.na(df_gender$gender2)] <- 4
df_gender$gender2 <- as.factor(df_gender$gender2)
levels(df_gender$gender2) <- c("men", "women", "no first name", "missing")df_ppf <- df_ppf %>%
left_join(df_gender)4.3 Time (career age)
df_ppf$time <- df_ppf$pub_year - df_ppf$phd_year
df_ppf$covid <- as.numeric(df_ppf$pub_year == 2020 | df_ppf$pub_year == 2021)
df_ppf$covid2020 <- as.numeric(df_ppf$pub_year == 2020)
df_ppf$covid2021 <- as.numeric(df_ppf$pub_year == 2021)
df_ppf$age25 <- as.numeric(df_ppf$phd_year > 2010)
df_ppf$age25_2 <- as.numeric(df_ppf$phd_year > 2015)
df_ppf$phd_year0 <- df_ppf$phd_year - 1990
df_ppf$stem <- ifelse((df_ppf$field == "Biological and Health Sciences" | df_ppf$field == "Physical and Mathematical Sciences" |
df_ppf$field == "Engineering" | df_ppf$field == "Agricultural Sciences"), "STEM-M", ifelse(df_ppf$field ==
"missing", "missing", "NON-STEM-M"))
df_ppf$ethnicity <- as.factor(df_ppf$ethnicity)
df_ppf$ethnicity <- relevel(df_ppf$ethnicity, 2)
# levels(df_ppf$ethnicity)
df_ppf$multidisc <- 1 - (df_ppf$health_domain^2 + df_ppf$life_domain^2 + df_ppf$physical_domain^2 + df_ppf$social_domain^2)5 Robustness checks
5.1 Model 2 separate per domain
5.1.1 Strict selection
I am currently making strict selections on domain. Namely, authors who only publish in a specific domain. A different strategy would be to use >.5 as criteria or classifying scholars based on highest score across the domains.
df3 <- df_ppf[, c("npubs", "npubs_a", "npubs_first_a", "npubs_last_a", "time", "id", "covid", "covid2020",
"covid2021", "gender2", "phd_year0", "ethnicity", "uni", "health_domain", "life_domain", "physical_domain",
"social_domain")]
df3 <- na.omit(df3)
knots5 <- quantile(df3$time, probs = seq(0, 1, 0.25))[2:4]
con <- gamlss.control(n.cyc = 150)
df3_d1 <- df3[df3$health_domain == 1, ]
df3_d2 <- df3[df3$life_domain == 1, ]
df3_d3 <- df3[df3$physical_domain == 1, ]
df3_d4 <- df3[df3$social_domain == 1, ]
m2_d1 <- gamlss(npubs ~ bSpline(time, knots = knots5, df = 3) + phd_year0 + covid + gender2 + ethnicity +
uni + covid * gender2, data = df3_d1, family = BNB, control = con)
fsave(m2_d1, "m2_d1")
sm2_d1 <- summary(m2_d1)
fsave(sm2_d1, "sm2_d1")
m2_d2 <- gamlss(npubs ~ bSpline(time, knots = knots5, df = 3) + phd_year0 + covid + gender2 + ethnicity +
uni + covid * gender2, data = df3_d2, family = BNB, control = con)
fsave(m2_d2, "m2_d2")
sm2_d2 <- summary(m2_d2)
fsave(sm2_d2, "sm2_d2")
m2_d3 <- gamlss(npubs ~ bSpline(time, knots = knots5, df = 3) + phd_year0 + covid + gender2 + ethnicity +
uni + covid * gender2, data = df3_d3, family = BNB, control = con)
fsave(m2_d3, "m2_d3")
sm2_d3 <- summary(m2_d3)
fsave(sm2_d3, "sm2_d3")
m2_d4 <- gamlss(npubs ~ bSpline(time, knots = knots5, df = 3) + phd_year0 + covid + gender2 + ethnicity +
uni + covid * gender2, data = df3_d4, family = BNB, control = con)
fsave(m2_d4, "m2_d4")
sm2_d4 <- summary(m2_d4)
fsave(sm2_d4, "sm2_d4")names <- c("Intercept: mu", "bSpline(time, knots = knots5, df = 3)1", "bSpline(time, knots = knots5, df = 3)2",
"bSpline(time, knots = knots5, df = 3)3", "bSpline(time, knots = knots5, df = 3)4", "bSpline(time, knots = knots5, df = 3)5",
"bSpline(time, knots = knots5, df = 3)6", "PhD-cohort", "COVID-19 years", "gender: women", "gender: no first name",
"gender: missing", "ethnicity: minority I", "ethnicity: minority II", "ethnicity: missing", "university: LU",
"university: RU", "university: RUG", "university: TUE", "university: TI", "university: UM", "university: UT",
"university: UU", "university: UvA", "university: VU", "university: WUR", "COVID*women", "COVID*no first name",
"COVID*missing", "Intercept: sigma", "Intercept: nu")
rownames(sm2_d1) <- names
names <- c("Intercept: mu", "bSpline(time, knots = knots5, df = 3)1", "bSpline(time, knots = knots5, df = 3)2",
"bSpline(time, knots = knots5, df = 3)3", "bSpline(time, knots = knots5, df = 3)4", "bSpline(time, knots = knots5, df = 3)5",
"bSpline(time, knots = knots5, df = 3)6", "PhD-cohort", "COVID-19 years", "gender: women", "gender: no first name",
"gender: missing", "ethnicity: minority I", "ethnicity: minority II", "ethnicity: missing", "university: LU",
"university: RU", "university: RUG", "university: UM", "university: UT", "university: UU", "university: UvA",
"university: VU", "university: WUR", "COVID*women", "COVID*no first name", "COVID*missing", "Intercept: sigma",
"Intercept: nu")
rownames(sm2_d2) <- names
names <- c("Intercept: mu", "bSpline(time, knots = knots5, df = 3)1", "bSpline(time, knots = knots5, df = 3)2",
"bSpline(time, knots = knots5, df = 3)3", "bSpline(time, knots = knots5, df = 3)4", "bSpline(time, knots = knots5, df = 3)5",
"bSpline(time, knots = knots5, df = 3)6", "PhD-cohort", "COVID-19 years", "gender: women", "gender: no first name",
"gender: missing", "ethnicity: minority I", "ethnicity: minority II", "ethnicity: missing", "university: LU",
"university: RU", "university: RUG", "university: TUD", "university: TUE", "university: UM", "university: UT",
"university: UU", "university: UvA", "university: VU", "university: WUR", "COVID*women", "COVID*no first name",
"COVID*missing", "Intercept: sigma", "Intercept: nu")
rownames(sm2_d3) <- names
names <- c("Intercept: mu", "bSpline(time, knots = knots5, df = 3)1", "bSpline(time, knots = knots5, df = 3)2",
"bSpline(time, knots = knots5, df = 3)3", "bSpline(time, knots = knots5, df = 3)4", "bSpline(time, knots = knots5, df = 3)5",
"bSpline(time, knots = knots5, df = 3)6", "PhD-cohort", "COVID-19 years", "gender: women", "gender: no first name",
"gender: missing", "ethnicity: minority I", "ethnicity: minority II", "ethnicity: missing", "university: LU",
"university: RU", "university: RUG", "university: TUD", "university: TUE", "university: TI", "university: UM",
"university: UT", "university: UU", "university: UvA", "university: VU", "university: WUR", "COVID*women",
"COVID*no first name", "COVID*missing", "Intercept: sigma", "Intercept: nu")
rownames(sm2_d4) <- names
fshowdf(sm2_d1)
fshowdf(sm2_d2)
fshowdf(sm2_d3)
fshowdf(sm2_d4)| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| Intercept: mu | 1.550 | 0.081 | 19.219 | 0.000 |
| bSpline(time, knots = knots5, df = 3)1 | -0.733 | 0.110 | -6.681 | 0.000 |
| bSpline(time, knots = knots5, df = 3)2 | -0.894 | 0.093 | -9.604 | 0.000 |
| bSpline(time, knots = knots5, df = 3)3 | -1.150 | 0.096 | -12.033 | 0.000 |
| bSpline(time, knots = knots5, df = 3)4 | -1.094 | 0.172 | -6.372 | 0.000 |
| bSpline(time, knots = knots5, df = 3)5 | -1.004 | 0.284 | -3.539 | 0.000 |
| bSpline(time, knots = knots5, df = 3)6 | -2.475 | 0.385 | -6.427 | 0.000 |
| PhD-cohort | -0.033 | 0.003 | -10.614 | 0.000 |
| COVID-19 years | -0.485 | 0.080 | -6.089 | 0.000 |
| gender: women | -0.240 | 0.036 | -6.722 | 0.000 |
| gender: no first name | 0.835 | 0.174 | 4.812 | 0.000 |
| gender: missing | 0.411 | 0.405 | 1.016 | 0.310 |
| ethnicity: minority I | 0.094 | 0.159 | 0.589 | 0.556 |
| ethnicity: minority II | 0.016 | 0.068 | 0.241 | 0.810 |
| ethnicity: missing | -0.242 | 0.066 | -3.660 | 0.000 |
| university: LU | -0.146 | 0.354 | -0.412 | 0.680 |
| university: RU | 0.278 | 0.045 | 6.205 | 0.000 |
| university: RUG | 0.382 | 0.152 | 2.519 | 0.012 |
| university: TUE | -1.607 | 0.461 | -3.482 | 0.000 |
| university: TI | 0.629 | 0.174 | 3.619 | 0.000 |
| university: UM | 0.309 | 0.056 | 5.519 | 0.000 |
| university: UT | -0.299 | 0.103 | -2.897 | 0.004 |
| university: UU | 0.273 | 0.062 | 4.427 | 0.000 |
| university: UvA | 0.296 | 0.079 | 3.733 | 0.000 |
| university: VU | 0.391 | 0.095 | 4.130 | 0.000 |
| university: WUR | 0.666 | 0.089 | 7.506 | 0.000 |
| COVID*women | -0.125 | 0.109 | -1.142 | 0.254 |
| COVID*no first name | 0.531 | 0.569 | 0.934 | 0.350 |
| COVID*missing | 0.026 | 0.809 | 0.032 | 0.974 |
| Intercept: sigma | -1.642 | 0.093 | -17.620 | 0.000 |
| Intercept: nu | -1.426 | 0.175 | -8.161 | 0.000 |
| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| Intercept: mu | 1.711 | 0.145 | 11.783 | 0.000 |
| bSpline(time, knots = knots5, df = 3)1 | -0.782 | 0.171 | -4.568 | 0.000 |
| bSpline(time, knots = knots5, df = 3)2 | -0.879 | 0.127 | -6.896 | 0.000 |
| bSpline(time, knots = knots5, df = 3)3 | -1.184 | 0.138 | -8.580 | 0.000 |
| bSpline(time, knots = knots5, df = 3)4 | -0.757 | 0.199 | -3.801 | 0.000 |
| bSpline(time, knots = knots5, df = 3)5 | -1.305 | 0.258 | -5.051 | 0.000 |
| bSpline(time, knots = knots5, df = 3)6 | -1.509 | 0.260 | -5.796 | 0.000 |
| PhD-cohort | -0.058 | 0.004 | -14.754 | 0.000 |
| COVID-19 years | -0.571 | 0.083 | -6.863 | 0.000 |
| gender: women | -0.096 | 0.049 | -1.972 | 0.049 |
| gender: no first name | -0.649 | 0.337 | -1.927 | 0.054 |
| gender: missing | -0.608 | 1.061 | -0.573 | 0.567 |
| ethnicity: minority I | -0.412 | 0.237 | -1.738 | 0.082 |
| ethnicity: minority II | -0.260 | 0.079 | -3.299 | 0.001 |
| ethnicity: missing | -0.271 | 0.059 | -4.564 | 0.000 |
| university: LU | 0.031 | 0.256 | 0.121 | 0.904 |
| university: RU | 0.129 | 0.103 | 1.246 | 0.213 |
| university: RUG | 1.013 | 0.172 | 5.892 | 0.000 |
| university: UM | 0.018 | 0.129 | 0.140 | 0.889 |
| university: UT | -0.471 | 0.351 | -1.342 | 0.180 |
| university: UU | -0.029 | 0.128 | -0.226 | 0.821 |
| university: UvA | 0.313 | 0.106 | 2.956 | 0.003 |
| university: VU | 0.391 | 0.125 | 3.115 | 0.002 |
| university: WUR | 0.588 | 0.093 | 6.355 | 0.000 |
| COVID*women | 0.200 | 0.120 | 1.662 | 0.097 |
| COVID*no first name | -0.359 | 0.851 | -0.422 | 0.673 |
| COVID*missing | -34.642 | 70710.678 | 0.000 | 1.000 |
| Intercept: sigma | -2.510 | 0.127 | -19.708 | 0.000 |
| Intercept: nu | -0.958 | 0.147 | -6.511 | 0.000 |
| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| Intercept: mu | 2.118 | 0.310 | 6.842 | 0.000 |
| bSpline(time, knots = knots5, df = 3)1 | -0.702 | 0.102 | -6.848 | 0.000 |
| bSpline(time, knots = knots5, df = 3)2 | -0.740 | 0.087 | -8.547 | 0.000 |
| bSpline(time, knots = knots5, df = 3)3 | -0.971 | 0.087 | -11.184 | 0.000 |
| bSpline(time, knots = knots5, df = 3)4 | -0.262 | 0.147 | -1.783 | 0.075 |
| bSpline(time, knots = knots5, df = 3)5 | -1.068 | 0.216 | -4.934 | 0.000 |
| bSpline(time, knots = knots5, df = 3)6 | -1.225 | 0.237 | -5.164 | 0.000 |
| PhD-cohort | -0.035 | 0.003 | -11.758 | 0.000 |
| COVID-19 years | -0.570 | 0.058 | -9.880 | 0.000 |
| gender: women | 0.039 | 0.041 | 0.957 | 0.339 |
| gender: no first name | -0.287 | 0.155 | -1.849 | 0.064 |
| gender: missing | 0.600 | 0.153 | 3.910 | 0.000 |
| ethnicity: minority I | -0.232 | 0.118 | -1.962 | 0.050 |
| ethnicity: minority II | -0.214 | 0.052 | -4.108 | 0.000 |
| ethnicity: missing | -0.026 | 0.039 | -0.674 | 0.500 |
| university: LU | 0.096 | 0.318 | 0.301 | 0.763 |
| university: RU | 0.188 | 0.304 | 0.619 | 0.536 |
| university: RUG | 0.685 | 0.331 | 2.066 | 0.039 |
| university: TUD | 0.594 | 0.317 | 1.878 | 0.060 |
| university: TUE | 0.459 | 0.303 | 1.518 | 0.129 |
| university: UM | -1.017 | 0.434 | -2.347 | 0.019 |
| university: UT | -0.236 | 0.302 | -0.781 | 0.435 |
| university: UU | 0.154 | 0.302 | 0.510 | 0.610 |
| university: UvA | 0.380 | 0.302 | 1.259 | 0.208 |
| university: VU | -0.012 | 0.305 | -0.040 | 0.968 |
| university: WUR | 0.014 | 0.303 | 0.047 | 0.963 |
| COVID*women | 0.135 | 0.109 | 1.245 | 0.213 |
| COVID*no first name | -0.355 | 0.738 | -0.481 | 0.630 |
| COVID*missing | -0.412 | 0.749 | -0.549 | 0.583 |
| Intercept: sigma | -0.376 | 0.049 | -7.695 | 0.000 |
| Intercept: nu | -0.788 | 0.133 | -5.926 | 0.000 |
| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| Intercept: mu | 1.232 | 0.105 | 11.711 | 0.000 |
| bSpline(time, knots = knots5, df = 3)1 | -0.438 | 0.079 | -5.560 | 0.000 |
| bSpline(time, knots = knots5, df = 3)2 | -0.478 | 0.063 | -7.597 | 0.000 |
| bSpline(time, knots = knots5, df = 3)3 | -0.670 | 0.063 | -10.644 | 0.000 |
| bSpline(time, knots = knots5, df = 3)4 | -1.006 | 0.108 | -9.331 | 0.000 |
| bSpline(time, knots = knots5, df = 3)5 | -1.125 | 0.180 | -6.256 | 0.000 |
| bSpline(time, knots = knots5, df = 3)6 | -2.696 | 0.255 | -10.580 | 0.000 |
| PhD-cohort | -0.044 | 0.002 | -20.955 | 0.000 |
| COVID-19 years | -0.571 | 0.056 | -10.196 | 0.000 |
| gender: women | -0.088 | 0.023 | -3.814 | 0.000 |
| gender: no first name | -0.327 | 0.107 | -3.050 | 0.002 |
| gender: missing | -0.146 | 0.295 | -0.494 | 0.621 |
| ethnicity: minority I | -0.103 | 0.104 | -0.991 | 0.322 |
| ethnicity: minority II | -0.184 | 0.046 | -3.982 | 0.000 |
| ethnicity: missing | -0.031 | 0.032 | -0.949 | 0.343 |
| university: LU | 0.592 | 0.115 | 5.162 | 0.000 |
| university: RU | 0.570 | 0.097 | 5.887 | 0.000 |
| university: RUG | 0.855 | 0.099 | 8.620 | 0.000 |
| university: TUD | 0.235 | 0.539 | 0.435 | 0.663 |
| university: TUE | 0.178 | 0.167 | 1.065 | 0.287 |
| university: TI | 0.695 | 0.117 | 5.930 | 0.000 |
| university: UM | 0.661 | 0.099 | 6.698 | 0.000 |
| university: UT | 0.554 | 0.105 | 5.270 | 0.000 |
| university: UU | 0.469 | 0.103 | 4.540 | 0.000 |
| university: UvA | 0.677 | 0.096 | 7.029 | 0.000 |
| university: VU | 0.632 | 0.098 | 6.426 | 0.000 |
| university: WUR | 0.467 | 0.145 | 3.225 | 0.001 |
| COVID*women | 0.060 | 0.078 | 0.762 | 0.446 |
| COVID*no first name | -0.190 | 0.401 | -0.473 | 0.636 |
| COVID*missing | 1.200 | 0.615 | 1.950 | 0.051 |
| Intercept: sigma | -1.887 | 0.080 | -23.494 | 0.000 |
| Intercept: nu | -1.643 | 0.185 | -8.871 | 0.000 |
5.2 Fixed effects career age and person id
df3 <- df_ppf[, c("npubs", "npubs_a", "npubs_first_a", "npubs_last_a", "time", "pub_year", "id", "covid",
"covid2020", "covid2021", "gender2", "phd_year0", "phd_year", "ethnicity", "uni", "life_domain",
"physical_domain", "social_domain")]
df3 <- na.omit(df3)
df3$log_npubs <- log(df3$npubs + 1)
m_fe <- feols(log_npubs ~ 1 + covid + i(gender2, covid, ref = c("men")) | id + time, data = df3)
fshowdf(summary(m_fe)$coeftable)| Estimate | Std. Error | t value | Pr(>|t|) | |
|---|---|---|---|---|
| covid | -0.284 | 0.009 | -33.260 | 0.000 |
| gender2::women:covid | 0.039 | 0.013 | 3.041 | 0.002 |
| gender2::no first name:covid | -0.097 | 0.059 | -1.643 | 0.100 |
| gender2::missing:covid | 0.026 | 0.103 | 0.249 | 0.804 |
Based on this modelling strategy we would conclude that women were significantly less negatively impacted by COVID-19 with respect to their absolute number of yearly articles.
5.3 Table 4
Staying close to modelling strategy of Madsen et al. (2022).
We do not use year of first pub to define career age but phd_year
df3 <- df_ppf[, c("npubs", "npubs_a", "npubs_first_a", "npubs_last_a", "time", "pub_year", "id", "covid",
"covid2020", "covid2021", "gender2", "phd_year", "phd_year0", "ethnicity", "uni", "life_domain",
"physical_domain", "social_domain")]
df3$log_npubs_a <- log(df3$npubs_a + 1)
df3_mid <- df3[df3$phd_year == 2008 | df3$phd_year == 2009 | df3$phd_year == 2010, ]
df3_early <- df3[df3$phd_year == 2014 | df3$phd_year == 2015 | df3$phd_year == 2016, ]
# remove missings on gender
df3_early <- df3_early[df3_early$gender2 == "men" | df3_early$gender2 == "women", ]
df3_mid <- df3_mid[df3_mid$gender2 == "men" | df3_mid$gender2 == "women", ]
df3_early$women <- ifelse(df3_early$gender2 == "women", 1, 0)
df3_mid$women <- ifelse(df3_mid$gender2 == "women", 1, 0)5.3.1 observations
print("N_persons (early career)")
length(unique(df3_early$id))
print("N_observations (early career)")
nrow(df3_early)
print("N_persons (mid career)")
length(unique(df3_mid$id))
print("N_observations (mid career)")
nrow(df3_mid)#> [1] "N_persons (early career)"
#> [1] 1310
#> [1] "N_observations (early career)"
#> [1] 8888
#> [1] "N_persons (mid career)"
#> [1] 1242
#> [1] "N_observations (mid career)"
#> [1] 14959m_fe_e <- feols(log_npubs_a ~ 1 + i(pub_year, women, ref = "2019") | id + pub_year, data = df3_early)
m_fe_m <- feols(log_npubs_a ~ 1 + i(pub_year, women, ref = "2019") | id + pub_year, data = df3_mid)5.3.2 fit measures
r2 <- c(fixest::r2(m_fe_e, "r2"), fixest::r2(m_fe_m, "r2")) #overal r2
wr2 <- c(fixest::r2(m_fe_e, "wr2"), fixest::r2(m_fe_m, "wr2")) #within r2
df_fitst4 <- data.frame(r2 = r2, wr2 = wr2)
# fsave(df_fitst4, 'df_fitst4')smfee <- as.data.frame(coeftable(m_fe_e))
smfee$names <- rownames(smfee)
smfem <- as.data.frame(coeftable(m_fe_m))
smfem$names <- rownames(smfem)
res <- right_join(smfee, smfem, by = "names")
res <- res[order(res$names), ]
res <- res[, -c(3, 8)]
res <- res[, c(4, 1:3, 5:7)]
res$names <- c("year2009*women", "year2010*women", "year2011*women", "year2012*women", "year2013*women",
"year2014*women", "year2015*women", "year2016*women", "year2017*women", "year2018*women", "year2020*women",
"year2021*women", "year2022*women")
colnames(res) <- as.character(1:7)
# fsave(res, 'Table4_raw')
Table4_raw <- restable4ft <- flextable(Table4_raw) %>% add_header_row(values=c(" ", rep(c("B", "S.E.", "P"), 2)), top=FALSE) %>% delete_rows(i=1,part="header") %>%
#set_caption( "Fixed-Effects model for yearly article output (logtransformed; Gaussian)") %>%
add_footer_lines(value = c("Notes: \nFor reasons of parsimony, the estimates of the fixed effects are not shown: scholars; year. Scholars with missing values on gender were removed from the analysis.\nEarly career: N_observations 8,888; N_persons 1,310 \n Mid career: N_obervations 14,959; N_persons: 1,242")) %>%
add_header_row(values = c(" ", "early career PhDs", "mid-career PhDs"), colwidths = c(1, 3, 3)) %>%
align(i = 1, align = "center", part = "header") %>%
align(i = 2, align = "center", part = "header") %>%
colformat_double(digits = 3) %>%
fontsize(size = 8, part = "all" ) %>%
set_table_properties(opts_pdf = list(arraystretch = 1)) %>% #to adjust row height
width(j = 1, width = 1.3) %>%
width(j = 2:7 , width = .4)
table4ft <- add_body_row(table4ft,
values = c("overall R-squared",round(df_fitst4$r2,3)),
colwidths = c(1,rep(3, 2)), top = FALSE
) %>%
align(i = 14, j=c(2:7), align = "center", part = "body")
table4ft <- add_body_row(table4ft,
values = c("within R-squared", round(df_fitst4$wr2,3)),
colwidths = c(1,rep(3, 2)), top = FALSE
) %>%
align(i = 15, j=c(2:7), align = "center", part = "body")
table4ft %>%
border_inner_h(border = fp_border_default(width = 0), part = "body") %>%
hline(i = 13)
| early career PhDs | mid-career PhDs | ||||
|---|---|---|---|---|---|---|
| B | S.E. | P | B | S.E. | P |
year2009*women | -0.066 | 0.080 | 0.411 | |||
year2010*women | -0.062 | 0.067 | 0.354 | |||
year2011*women | -0.074 | 0.057 | 0.194 | |||
year2012*women | -0.040 | 0.053 | 0.454 | |||
year2013*women | -0.041 | 0.049 | 0.404 | |||
year2014*women | -0.039 | 0.049 | 0.431 | |||
year2015*women | 0.071 | 0.065 | 0.276 | -0.009 | 0.045 | 0.836 |
year2016*women | 0.084 | 0.046 | 0.070 | -0.032 | 0.042 | 0.452 |
year2017*women | 0.033 | 0.039 | 0.394 | -0.027 | 0.041 | 0.514 |
year2018*women | 0.007 | 0.037 | 0.858 | -0.018 | 0.037 | 0.637 |
year2020*women | 0.003 | 0.038 | 0.935 | 0.007 | 0.039 | 0.852 |
year2021*women | 0.025 | 0.039 | 0.529 | 0.074 | 0.043 | 0.086 |
year2022*women | 0.085 | 0.039 | 0.030 | 0.041 | 0.044 | 0.349 |
overall R-squared | 0.53 | 0.524 | ||||
within R-squared | 0.001 | 0.001 | ||||
Notes: | ||||||
Based on this modelling strategy we conclude that the gender gaps in publication were not more severe for women during COVID-19 than in pre- or post-COVID-19 years.
5.3.3 ATT
Dashed line is counterfactual. The difference between the blue dashed line and the solid dashed line is what (Madsen et al. 2022) interpret as the ATT.
5.3.3.1 Early career
df3_early$preds <- exp(predict(m_fe_e) - 1)
df3_early_new <- df3_early
df3_early_new <- df3_early_new %>%
mutate(women = case_when(pub_year > 2018 ~ 0, .default = women))
df3_early_new$preds2 <- exp(predict(m_fe_e, newdata = df3_early_new) - 1)
df3_early_new$women <- df3_early$women
df3_early_ag <- df3_early_new %>%
group_by(women, pub_year) %>%
summarise(yhat_mean = mean(preds), yhat_mean_c = mean(preds2))5.3.3.2 Mid career
df3_mid$preds <- exp(predict(m_fe_m) - 1)
df3_mid_new <- df3_mid
df3_mid_new <- df3_mid_new %>%
mutate(women = case_when(pub_year > 2018 ~ 0, .default = women))
df3_mid_new$preds2 <- exp(predict(m_fe_m, newdata = df3_mid_new) - 1)
df3_mid_new$women <- df3_mid$women
df3_mid_ag <- df3_mid_new %>%
group_by(women, pub_year) %>%
summarise(yhat_mean = mean(preds), yhat_mean_c = mean(preds2))gec <- ggplot(df3_early_ag, aes(x = pub_year, y = yhat_mean)) + geom_line(aes(color = as.factor(women)),
show.legend = FALSE) + geom_line(aes(y = yhat_mean_c, color = as.factor(women)), show.legend = FALSE,
linetype = "dashed") + ylim(0, 1.5) + xlab("year") + ylab("publication count") + theme_minimal() +
theme(plot.caption = element_text(hjust = 0)) + theme(plot.margin = margin(t = 1, r = 1, b = 1, l = 1,
unit = "cm"))
gmc <- ggplot(df3_mid_ag, aes(x = pub_year, y = yhat_mean)) + geom_line(aes(color = as.factor(women)),
show.legend = TRUE) + geom_line(aes(y = yhat_mean_c, color = as.factor(women)), show.legend = FALSE,
linetype = "dashed") + ylim(0, 1.5) + xlab("year") + ylab("publication count") + scale_colour_discrete(name = "Gender",
breaks = c("0", "1"), labels = c("men", "women")) + theme_minimal() + theme(plot.caption = element_text(hjust = 0)) +
theme(plot.margin = margin(t = 1, r = 1, b = 1, l = 1, unit = "cm"))
Figure_RR2 <- ggarrange(gec, gmc, ncol = 2, labels = c("early career", "mid career"), hjust = c(-0.5,
-0.5), nrow = 1, common.legend = TRUE, legend = "right")
Figure_RR2 <- annotate_figure(Figure_RR2, bottom = text_grob("This figure shows the (mean) predicted publishing rates for men and women authors, with solid lines showing \nthe trend per gender, and the dashed blue line showing the counterfactual trend for women if they had similar \n2019–2022 trajectories as men. The difference between the dashed blue line and the straight blue line is \ninterpreted by Madsen et al. (2022) as the average treatment effect for women.",
color = "black", just = "left", x = 0.05, y = 1, size = 10, lineheight = 1))
Figure_RR2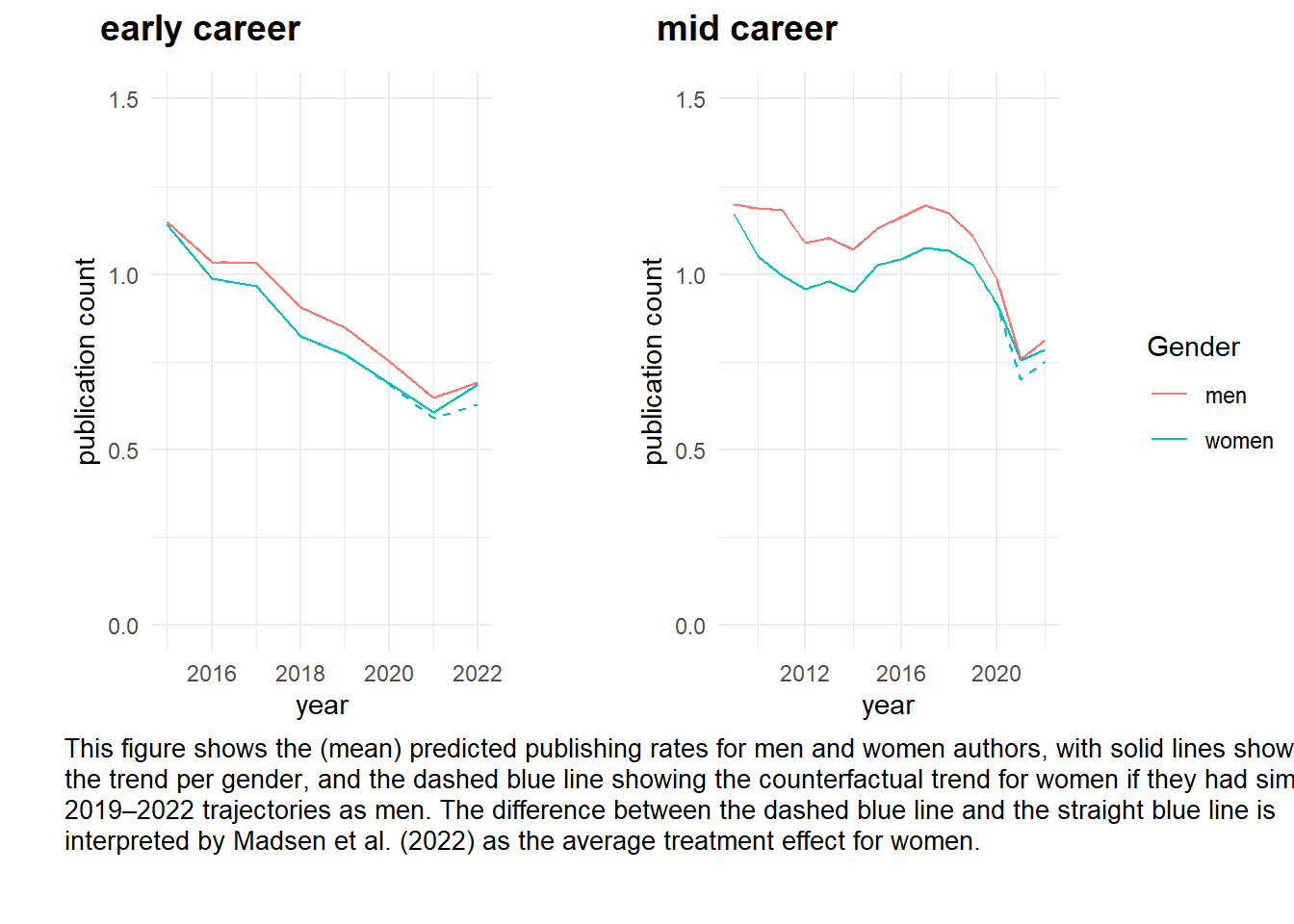
# now find a way to save correctly { pdf(file = './data/processed/20250729FigureRR2.pdf', width =
# 10, height =10) Figure_RR2 } dev.off() { png(file = './data/processed/20250729FigureRR2.png',
# width = 1600, height = 1600, units = 'px', res = 200) Figure_RR2 } dev.off()Based on these figure we conclude that the ‘ATT’ for women is negative, women were less (negatively) impacted by COVID-19 than men.
Copyright © 2024- Jochem Tolsma